May 1, 2020
Created Using Covid19Primer.com
Testing, Testing and More Testing
Random Testing the Population Could be the Key to Getting Out of a Lockdown Earlier
Swiss scientists have shown how randomly testing the population can allow for a reboot of the economy much quicker and more safely than waiting the 14-day period for symptoms to show. Using random testing in a feedback-control loop to manage a safe exit from the COVID-19 lockdown
The scientists showed that with daily sampling in place, a reboot of the economy “could be attempted while the fraction of infected people is still an order of magnitude higher than the level required for a relaxation of restrictions with testing focused on symptomatic individuals”. Meaning that with random testing in place, countries can potentially start getting back to normal much sooner.
It doesn’t have to be PCR testing either. The researchers state that “Any other measurement of the fraction of currently infected people can replace the random testing. For example, there are proposals to estimate this fraction from analysis of sewage water with PCR test.”
However, as Shenta et al said in their paper, “a major bottleneck of managing the COVID-19 pandemic in many countries is diagnostic testing, due to limited laboratory capabilities as well as limited access to genome-extraction and Polymerase Chain Reaction (PCR) reagents.”
But you don’t need to test every sample individually.
Less Testing, Better Results: Pooling the Tests Together
If you have 1000 people that you want to test for COVID-19, it turns out that testing each of them individually is not the most efficient way to do this. Scientists showed in this paper that for a prevalence of 10% of positive tests, you can reduce the number of tests needed by 40.6% by using a process that combines together the samples from groups of 4 people.
The scientists concluded that “Pool testing individuals for SARS‐CoV‐2 is a valuable strategy that could considerably boost a country’s testing capacity. However, further studies are needed to address how large these groups can be without losing sensitivity on the RT‐PCR.”
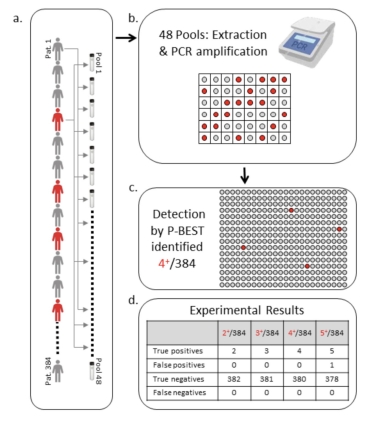
Researchers from Israel also showed similar results in their paper “Efficient high throughput SARS-CoV-2” where they developed a pooling method called P-BEST (Pooling-Based Efficient SARS-CoV-2 Testing).
“Instead of testing each sample separately, samples are pooled into groups and each pool is tested for SARS-CoV-2 using the standard clinically approved PCR-based diagnostic assay. Each sample is part of multiple pools, using a combinatorial pooling strategy based on compressed sensing designed for maximizing the ability to identify all positive individuals.”

They found that this method created an 8-fold increase in testing efficiency when compared with testing everyone’s individual sample.
“In our current proof-of-concept study we pooled 384 patient samples into 48 pools providing an 8-fold increase in testing efficiency.”
This research into the most efficient method to pool together samples is getting a lot of traction with multiple papers. Including this paper “Increased PCR screening capacity using a multi-replicate pooling scheme” where the scientists created free software to determine the optimal pooling strategy. The software can be downloaded here.
Testing the Wastewater to Predict Outbreaks
Taking pooling to another level, scientists have shown that they can not only test wastewater to estimate the prevalence of an outbreak in a population, they can do so on a massive scale. “We can actually measure hundreds of thousands of people with a single sample,” says Rolf Halden, the director of the Biodesign Center for Environmental Health Engineering at ASU.
Several research groups have reported detecting coronavirus in wastewater. Researchers from Paris, in their paper “Time course quantitative detection of SARS-CoV-2 in Parisian wastewaters correlates with COVID-19 confirmed cases” say they are the first to show that analysis and testing of wastewater can be predictive of future growth of the virus before these cases hit the clinics.
Science magazine reported on their work. “Sewer monitoring can illustrate the timing and scale of outbreaks that are currently difficult to visualize because of a general lack of human testing,” says Zhugen Yang, a biomedical engineer at Cranfield University’s Water Science Institute, a U.K. center that is developing $2 tests detecting SARS-CoV-2 in sewage. “Sewer sampling gives a fairly inexpensive, evidence-based image of the actual viral load in a community.”
Researchers from Spain have also found similar results. SARS-CoV-2 RNA titers in wastewater anticipated COVID-19 occurrence in a low prevalence area. “This environmental surveillance data were compared to declared COVID-19 cases at municipality level, revealing that SARS-CoV-2 was circulating among the population even before the first cases were reported by local or national authorities in many of the cities where wastewaters have been sampled. The detection of SARS-CoV-2 in wastewater in early stages of the spread of COVID-19 highlights the relevance of this strategy as an early indicator of the infection within a specific population.”
They also highlighted the immediate policy implications of this approach: “At this point, this environmental surveillance could be implemented by municipalities right away as a tool, designed to help authorities to coordinate the exit strategy to gradually lift its coronavirus lockdown.”
And it’s Much, Much Cheaper
Researchers from Tempe, AZ published similar findings for the US population in their paper: “Computational analysis of SARS-CoV-2/COVID-19 surveillance by wastewater-based epidemiology locally and globally: Feasibility, economy, opportunities and challenges.”
Popular Science reported on their work: “Researchers calculated that it should be possible to collect fecal data from the nation’s 15,014 wastewater treatment plants within several days. This screen would cover about 70 percent of the U.S. population.”
“The whole venture would cost about $225,000. By contrast, the team estimated, diagnostic tests for all 330 million or so Americans would cost around $3.5 billion. Sewage testing can never fully replace traditional diagnostic tests. But sewage sampling would make diagnostic testing more efficient by signaling where officials should be testing people more aggressively.”
Reading the Clinical Notes: All of Them
Researchers found that if they trained a computer to read through clinical notes from a large sample of patients they could start to predict who would be more likely to later test positive for COVID-19.
The research published on the MedXriv preprint server was in collaboration with the Mayo Clinic. The researchers analyzed “8.2 million clinical notes from 14,967 patients subjected to COVID-19 PCR diagnostic testing.” They found that these notes were predictive of COVID-19 positive tests. They were able to go further and identify which conditions were most predictive:
“We identify diarrhea (2.8-fold), change in appetite (2-fold), anosmia/dysgeusia (28.6-fold), and respiratory failure (2.1-fold) as significantly amplified in COVIDpos over COVIDneg patients.”

Combinations of these conditions were found to be more predictive: “The specific combination of cough and diarrhea has a 4-fold amplification in COVIDpos patients during the week prior to PCR testing, and along with anosmia/dysgeusia, constitutes the earliest EHR-derived signature of COVID-19 (4-7 days prior to typical PCR testing date).”
Other researchers from Mount Sinai have published work combining this type of clinical data analysis with image analysts from CT scans. The combination of the images and text analysis increased accuracy of their predictions. The researchers claim that “The proposed AI system achieved an AUC of 0.92 and performed equally well in sensitivity compared to a senior thoracic radiologist on a testing set of 279 cases.”
Can’t Test? Counting the Excess Deaths Might Provide Answers
In the research paper “COVID-19: a need for real-time monitoring of weekly excess deaths” published in the Lancet, the authors claimed that in order to measure the true extent of the pandemic that testing alone was insufficient and that excess deaths would provide a better measurement. The authors also claimed that “weekly excess deaths could provide the most objective and comparable way of assessing the scale of the pandemic and formulating lessons to be learned.”
Research into using excess deaths was also done by researchers from Italy. Their paper “Total COVID-19 Mortality in Italy: Excess Mortality and Age Dependence through Time-Series Analysis” was published on the preprint server MedRxiv. Modi et al claimed that “our analysis shows good agreement with reported COVID-19 mortality for age<70 years, but an excess in total mortality increasing with age above 70 years, suggesting there is a large population of predominantly old people missing from the official fatality statistics. We estimate that the number of COVID-19 deaths in Italy is 52,000 ± 2000 as of April 18 2020, more than a factor of 2 higher than the official number.”
More Testing Sites: A How to Guide
Scalable and Resilient SARS-CoV2 testing in an Academic Centre. “We provide a roadmap instructing how a research institute can be repurposed in the midst of this crisis, in collaboration with partner hospitals and an established diagnostic laboratory, harnessing existing expertise in virus handling, robotics, PCR, and data science to derive a rapid, high throughput diagnostic testing pipeline for detecting SARS-CoV-2 in patients with suspected COVID-19.”

“This strategy facilitates the remote reporting of thousands of samples a day with a turnaround time of under 24 hours, universally applicable to laboratories worldwide.”
Matt Buchanan tweeted: “London-based academic research center has repurposed resources to build an end-to-end pipeline for COVID-19 diagnostic testing 1000s tests/day, results in 24 hours Their new paper is a how-to guide for other institutions to follow”
Not to be outdone, researchers from Berkeley led by Jennifer Doudna published this paper: “Blueprint for a Pop-up SARS-CoV-2 Testing Lab.”

It was met with a positive response online with Alexander Ehrenburg tweeting: “How do you set up a fully functioning, certified, and FDA-compliant clinical lab from scratch in 3 wks? See this how-to. Obviously doesn’t hurt to have a rockstar team of leaders.”
Setting up pop-up labs and converting academic labs into testing facilities is one thing. But what about the commercial labs? It turns out from this survey in the journal Nature that more US labs do want to help – but can’t.

“Of the more than 4,000 researchers who responded within the first week, about 130 were already running tests to detect the new coronavirus. Nearly 1,600 said that they had the main tool needed to run tests, a real-time PCR machine, and operated under the biosafety conditions required for working with pathogenic organisms such as coronavirus. But they were not testing.”

False Negative in RT-PCR Tests
FALSE-NEGATIVE RESULTS OF INITIAL RT-PCR ASSAYS FOR COVID-19: A SYSTEMATIC REVIEW. “[O]ur findings reinforce the need for repeated testing in patients with suspicion of SARS-Cov-2 infection given that up to 29% of patients could have an initial RT-PCR false-negative result.”
RT-PCR Testing With Fewer Steps
Preliminary support for a ‘dry swab, extraction free’ protocol for SARS-CoV-2 testing via RT-qPCR. “As testing has scaled across the world, supply chain challenges have emerged across this entire workflow. […] Here we sought to evaluate how eliminating the UTM storage and RNA extraction steps would impact the results of molecular testing. […]We performed a comparison of conventional (swab → UTM → RNA extraction → RT-qPCR) vs. simplified (direct elution from dry swab → RT-qPCR) protocols. Our results suggest that dry swabs eluted directly into a simple buffered solution (TE) can support molecular detection of SARS-CoV-2 via endpoint RT-qPCR without substantially compromising sensitivity.”
Alternatives to RT-PCR Testing Methods
The majority of COVID-19 diagnostic tests rely on a technique called real-time reverse transcription-polymerase chain reaction (RT-PCR). This test amplifies SARS-CoV-2 RNA from patient swabs, but can take more than three hours to return a result as the viral RNA has to be prepared for analysis.
There are alternative testing methods being proposed including this research from a team out of Korea: “Rapid Detection of COVID-19 Causative Virus (SARS-CoV-2) in Human Nasopharyngeal Swab Specimens Using Field-Effect Transistor-Based Biosensor.”

The press release from EurekAlert is here. “Researchers reporting in ACS Nano have developed a field-effect transistor-based biosensor that detects SARS-CoV-2 in nasopharyngeal swabs from patients with COVID-19, in less than one minute.”
“The team based their test on a field-effect transistor — a sheet of graphene with high electronic conductivity. The researchers attached antibodies against the SARS-CoV-2 spike protein to the graphene. When they added either purified spike protein or cultured SARS-CoV-2 virus to the sensor, binding to the antibody caused a change in the electrical current. Next, the team tested the technique on nasopharyngeal swabs collected from patients with COVID-19 or healthy controls. Without any sample preparation, the sensor could discriminate between samples from sick and healthy patients.”
The test however, is not currently as accurate as the RT-PCR common tests. With the current materials researchers estimate that it is about 2-4 times less sensitive than RT-PCR, “…but different materials could be explored to improve the signal-to-noise ratio, the researchers say.”
Turning on the Lamp
Researchers from Israel “developed a protocol based on Reverse Transcribed Loop-Mediated Isothermal Amplification (RT-LAMP) for detection of SARS-CoV-2, directly from crude nose and throat swabs.”
The researchers claim that one of the benefits of this method is that “the proposed protocol provides results on-the-spot, and can detect SARS-CoV-2 from saliva”. You can see below a schematic of the test from their paper.

Another preprint paper from a research team from the Mason lab at Weill Cornell found similar results for LAMP testing. Here the researchers “designed a fast (30 minute) colorimetric test to identify SARS- CoV-2 infection”.
Using LAMP for COVID-19 testing is “a great idea that just needs to be tweaked and optimized,” noted Chris Mason, PhD, associate professor at Weill Cornell Medical College and senior author on the paper.
The research was covered in this in-depth article from Genetic Engineering and Biotechnology News which had this good explainer about the process:
“LAMP relies on DNA polymerase and a set of four to six primers, designed to recognize a total of six distinct sequences on the target DNA. One of the inner primers initiates strand displacement and amplification. A strand displacing DNA polymerase displaces the two original DNA strands while creating a new strand. A separate primer then displaces the newly made strand, and a loop is formed due to complementary sequences on the primers. The process of displacement, amplification, and forming loops continues until the DNA forms a dumbbell structure. It is the accumulation of these amplification byproducts that can be easily detected.”
If you still want more insight, here’s a bonus tutorial video explaining LAMP.
Saliva Seems to be a Good Substitute for Nasal Swabs
Saliva is more sensitive for SARS-CoV-2 detection in COVID-19 patients than nasopharyngeal swabs. “The current gold standard for COVID-19 diagnosis is real-time RT-PCR detection of SARS-CoV-2 from nasopharyngeal swabs. Low sensitivity, exposure risks to healthcare workers, and global shortages of swabs and personal protective equipment, however, necessitate the validation of new diagnostic approaches.”
The researchers from Yale found that “When we compared SARS-CoV-2 detection from patient-matched nasopharyngeal and saliva samples, we found that saliva yielded greater detection sensitivity and consistency throughout the course of infection. Furthermore, we report less variability in self-sample collection of saliva.”
The research may have significant testing policy implications and could allow widespread testing at home. The researchers claim that “Taken together, our findings demonstrate that saliva is a viable and more sensitive alternative to nasopharyngeal swabs and could enable at-home self-administered sample collection for accurate large-scale SARS-CoV-2 testing.”
The researchers also found instances where the saliva tests worked when the nasal tests didn’t. “Moreover, the detection of SARS-CoV-2 from the saliva of two asymptomatic healthcare workers despite negative matched nasopharyngeal swabs suggests that saliva may also be a viable alternative for identifying mild or subclinical infections.”
Antibody Testing
While PCR testing can detect if you’ve been exposed to the virus, only antibody testing can determine whether you’ve developed immunity. False positives in an antibody test can put people in harm’s way, as it might lead to a lifting of lockdowns before enough of the population has developed potential immunity.
The UK National COVID-19 Testing Scientific Advisory Panel is weighing in on this: “Evaluation of antibody testing for SARS-CoV-2 using ELISA and lateral flow immunoassays.”
“If antibody tests were deployed as an individual-level approach to inform release from quarantine, then high specificity is essential, as false-positive results return non-immune individuals to risk of exposure. For this reason, the UK Medicines and Healthcare products Regulatory Agency has set a minimum 98% specificity threshold for LFIAs. […] The early promise of lateral flow immunoassay (LFIA) devices has been questioned following concerns about sensitivity and specificity.”
“We tested plasma for SARS-Cov-2 IgM and IgG antibodies by ELISA and using nine different commercially available LFIA devices. […] Point estimates for the sensitivity of LFIA devices ranged from 55-70% versus RT-PCR and 65-85% versus ELISA, with specificity 95-100% and 93-100% respectively.”

Meta Analysis of Different Antibody Tests
Antibody tests in detecting SARS-CoV-2 infection: a meta-analysis. Researchers scanned all of the published research and “identified 38 eligible studies that include data from 7,848 individuals.”
“We evaluated IgM and IgG tests based on Enzyme-linked immunosorbent assay (ELISA), Chemiluminescence Enzyme Immunoassays (CLIA), Fluorescence Immunoassays (FIA) and the point-of-care (POC) Lateral Flow Immunoassays (LFIA) that are based on immunochromatography. […] IgG tests perform better compared to IgM ones, and show better sensitivity when the samples were taken longer after the onset of symptoms. […] ELISA- and CLIA-based methods performed better in terms of sensitivity (90-94%) followed by LFIA and FIA with sensitivities ranging from 80% to 86%. ELISA tests could be a safer choice at this stage of the pandemic.”
Here is a solid background briefing on antibody testing to help you get up to speed: Q&A on COVID-19 Antibody Tests from factcheck.org.
When it Comes to Antibodies: No Testing is Better Than Bad Testing?
“No test is better than a bad test”: Impact of diagnostic uncertainty in mass testing on the spread of Covid-19. “Diagnostic uncertainty can have a large effect on the epidemic dynamics of Covid-19 within the UK. The dynamics of the epidemic are more sensitive to test performance and targeting than test capacity. The quantity of tests is not a substitute for an effective strategy. Poorly targeted testing has the propensity to exacerbate the peak in infections. Interpretation: The assessment that ‘no test is better than a bad test’ is broadly supported by the present analysis. Antibody testing is unlikely to be a solution to the lock-down, regardless of test quality or capacity. A well designed active viral testing strategy combined with incremental relaxation of the lock-down measures is shown to be a potential strategy to restore some social activity whilst continuing to keep infections low.”
The research is very well covered in this article where you can read more about the policy implications of this study.
Continuous Temperature Monitoring
Continuous temperature monitoring by a wearable device for early detection of febrile events in the SARS-CoV-2 outbreak in Taiwan, 2020. “The HEARThermo, a watch-like wearable device, can measure body surface temperature and heart rate every 10s with good test-retest reliability and adequate criterion validity.” It can continuously monitor a person’s temperature to see if there are significant increases over time which may be indicative of COVID-19. Below is a sample graph from the paper of a subject’s temperature over time.

Some Good News
All essential workers in England will now be able to get tested. The British Medical Journal noted: “All key workers in England and members of their households who are showing symptoms of COVID-19 will now be able to get tested for the virus, the government has promised. This widening of access, enabled by an increase in capacity, comes as the government is under pressure to meet its target to carry out 100 000 tests a day by the end of April.”